top of page
Human Development & Aging
Gene expression in development (transcription factors, enhancers, silencers)
Core Principle of Developmental Gene Expression
🧷
Development transforms a single fertilized egg into a complex organism through precisely orchestrated gene expression programs.
🧷
Key molecular players include transcription factors (proteins that bind DNA and regulate transcription), enhancers (DNA sequences that increase transcription when bound by activators), and silencers (DNA sequences that decrease transcription when bound by repressors).
🧷
The fundamental mechanism: transcription factors bind to regulatory DNA elements → recruit cofactors → modify chromatin structure and RNA polymerase activity → control when, where, and how much each gene is expressed.
🧷
This spatiotemporal control creates distinct cell types from identical genomes.

Transcription Factors: The Master Regulators
📍
Transcription factors (TFs) are proteins containing DNA-binding domains that recognize specific 6-10 base pair sequences called motifs.
📍
Common DNA-binding domains include helix-turn-helix, zinc fingers, leucine zippers, and helix-loop-helix structures.
📍
TFs function as activators (increase transcription) or repressors (decrease transcription) by recruiting coactivators or corepressors.
📍
Many developmental TFs work in combinatorial codes — different combinations of TFs in a cell determine its fate.
📍
Board pearl: Mutations in transcription factor genes often cause developmental syndromes affecting multiple organ systems because one TF controls many target genes.

Enhancers: Remote Control Elements
🔹
Enhancers are regulatory DNA sequences that increase transcription from distances up to 1 million base pairs away from their target promoter.
🔹
They work regardless of orientation (forward or reverse) and position (upstream, downstream, or within introns) relative to the gene.
🔹
Mechanism: TFs bind enhancers → DNA loops bring enhancer-bound factors to the promoter → increased RNA polymerase II recruitment and activation.
🔹
A single gene often has multiple enhancers that drive expression in different tissues or developmental stages.
🔹
Board pearl: Enhancer mutations can cause disease even when the coding sequence is normal — they disrupt tissue-specific gene expression patterns.

Silencers and Repression Mechanisms
⭐
Silencers are DNA sequences that decrease transcription when bound by repressor proteins.
⭐
Repression mechanisms include: blocking activator binding sites, recruiting histone deacetylases (HDACs) to compact chromatin, recruiting DNA methyltransferases, and directly inhibiting RNA polymerase.
⭐
Polycomb group proteins are key developmental repressors that maintain gene silencing through histone H3K27 trimethylation.
⭐
Insulator elements prevent inappropriate enhancer-promoter interactions, maintaining proper gene expression boundaries.
⭐
Board distinction: Enhancers increase transcription regardless of position; silencers decrease it. Both can act at great distances from the promoter.

Homeobox Genes and HOX Clusters
✅
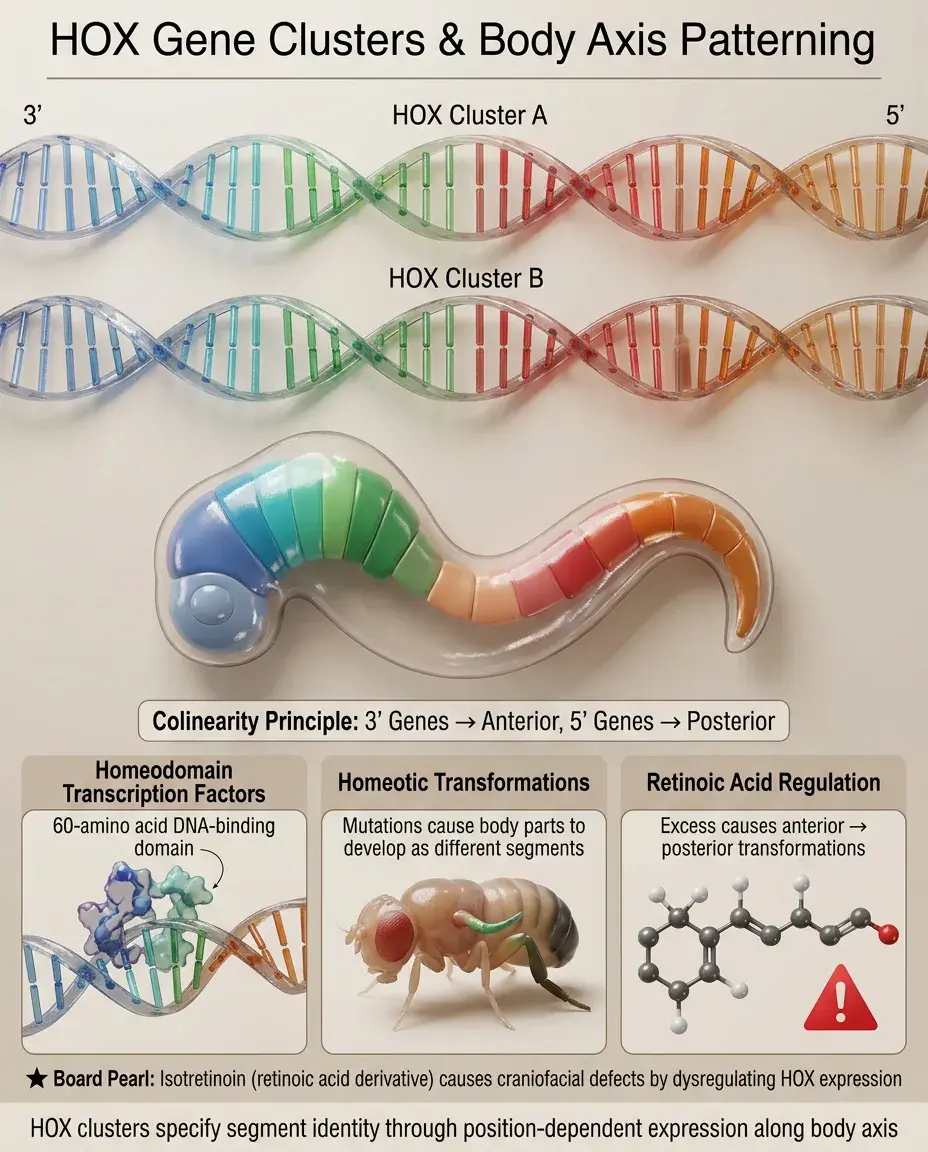
Homeobox genes encode transcription factors containing a 60-amino acid homeodomain that binds DNA.
✅
HOX genes are homeobox genes arranged in four clusters (A-D) on different chromosomes that control anterior-posterior body axis patterning.
✅
Colinearity principle: HOX gene position on the chromosome corresponds to expression along the body axis — 3' genes expressed anteriorly, 5' genes posteriorly.
✅
HOX proteins specify segment identity: mutations cause homeotic transformations (one body part develops as another).
✅
Board pearl: Retinoic acid regulates HOX gene expression — excess causes anterior structures to adopt posterior fates (e.g., isotretinoin causing craniofacial defects).
Master Regulatory Genes and Cell Fate
🧠
Master regulatory genes encode transcription factors sufficient to drive entire developmental programs.
🧠
PAX6: master regulator of eye development. Ectopic expression can induce eye formation in abnormal locations.
🧠
MyoD: master regulator of muscle. Can convert fibroblasts to muscle cells when artificially expressed.
🧠
SOX9: master regulator of male sex determination. Mutations cause campomelic dysplasia with sex reversal.
🧠
Board pearl: Master regulatory genes sit atop hierarchical cascades — one TF activates many downstream TFs, which activate even more, amplifying and refining the developmental program.

Morphogen Gradients and Transcriptional Responses
⚡
Morphogens are signaling molecules that form concentration gradients and elicit different cell fates at different concentrations.
⚡
Classic morphogens include Sonic hedgehog (SHH), BMPs, Wnts, and retinoic acid.
⚡
Cells interpret morphogen concentrations through dose-dependent activation of different target genes via threshold-sensitive enhancers.
⚡
Example: In the neural tube, high SHH (ventral) induces floor plate, medium SHH induces motor neurons, low SHH induces interneurons.
⚡
Board pearl: Holoprosencephaly results from SHH pathway mutations — failure to pattern the ventral forebrain causes midline defects (cyclopia, single central incisor).

Epigenetic Regulation in Development
📌
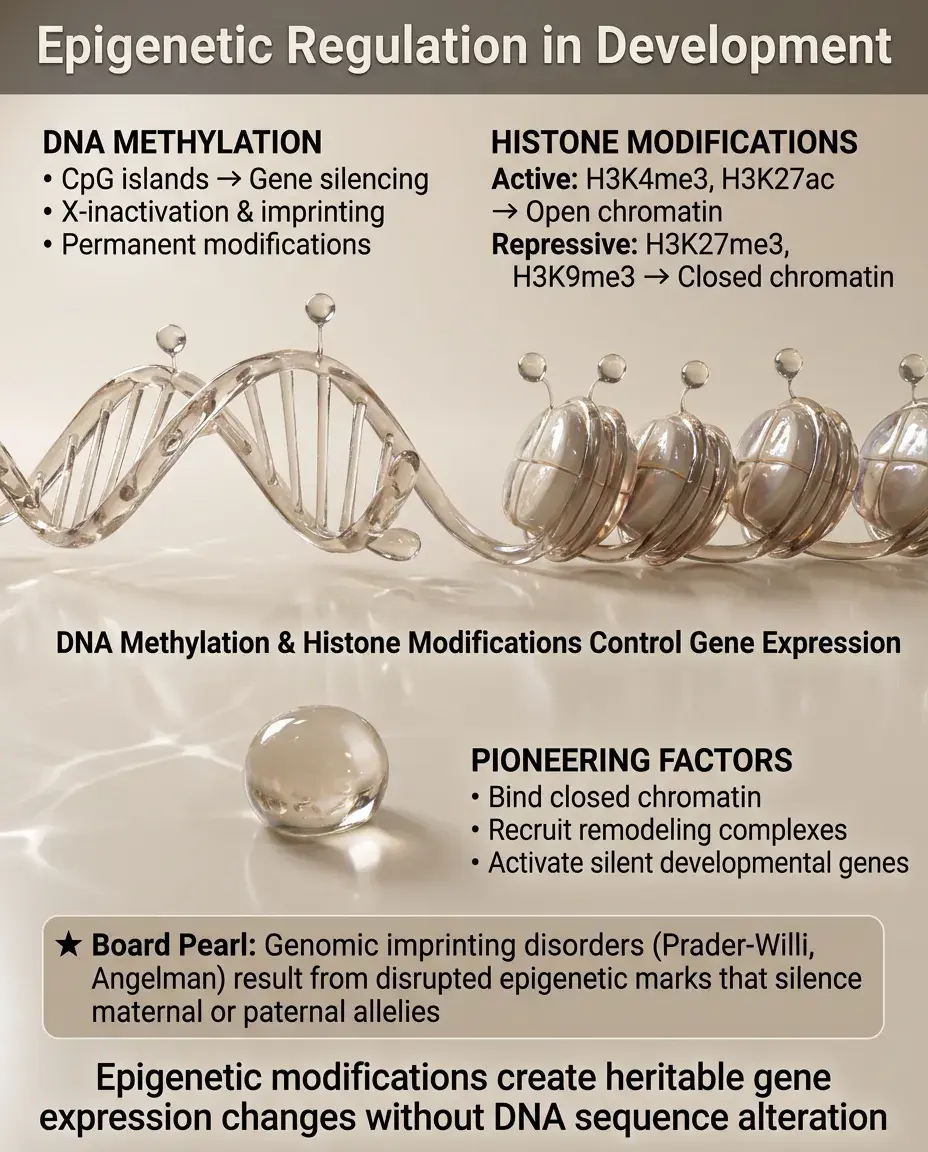
Epigenetic modifications — DNA methylation and histone modifications — create heritable changes in gene expression without altering DNA sequence.
📌
DNA methylation at CpG islands silences genes, often permanently (e.g., X-inactivation, imprinted genes).
📌
Histone modifications create active (H3K4me3, H3K27ac) or repressive (H3K27me3, H3K9me3) chromatin states.
📌
Pioneering transcription factors can bind to closed chromatin and recruit chromatin remodelers to activate silent genes.
📌
Board pearl: Genomic imprinting disorders (Prader-Willi, Angelman) result from disrupted epigenetic marks that normally silence either the maternal or paternal allele.
Combinatorial Control and Cell Type Specification
📣
Cell fate is determined by the combination of transcription factors expressed, not any single factor — this is combinatorial control.
📣
Example: Pancreatic β-cells require PDX1 + NKX6.1 + MAFA; changing this combination produces different cell types.
📣
Cross-repressive interactions between TFs create bistable switches — once a cell chooses a fate, it actively represses alternative fates.
📣
Feed-forward loops (A activates B, A+B together activate C) lock in developmental decisions and make them irreversible.
📣
Board pearl: Induced pluripotent stem cells (iPSCs) demonstrate that the right TF combination (Oct4, Sox2, Klf4, c-Myc) can override established cell fate.

Signal Transduction to Transcriptional Output
🔸
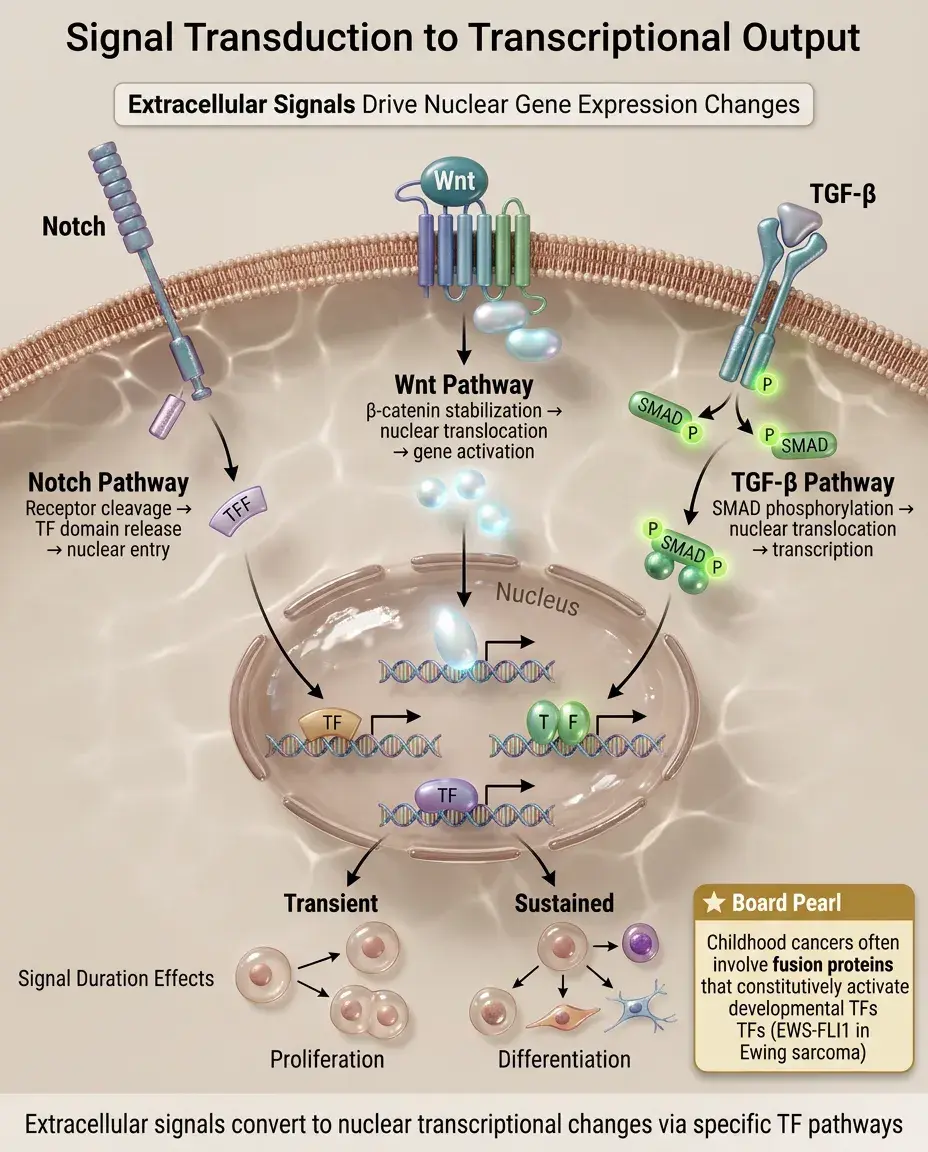
Extracellular signals (growth factors, morphogens) must be converted to transcriptional changes inside the nucleus.
🔸
Common pathway: ligand → receptor → signal transduction cascade → activation or nuclear translocation of transcription factors → gene expression changes.
🔸
Examples: Notch pathway (receptor cleavage releases TF domain), Wnt pathway (β-catenin stabilization and nuclear entry), TGF-β pathway (SMAD phosphorylation and nuclear translocation).
🔸
Signal duration matters: transient signals often induce proliferation, sustained signals induce differentiation.
🔸
Board pearl: Many childhood cancers involve translocation-created fusion proteins that constitutively activate developmental TFs (e.g., EWS-FLI1 in Ewing sarcoma).
Developmental Enhancers and Tissue Specificity
🧷
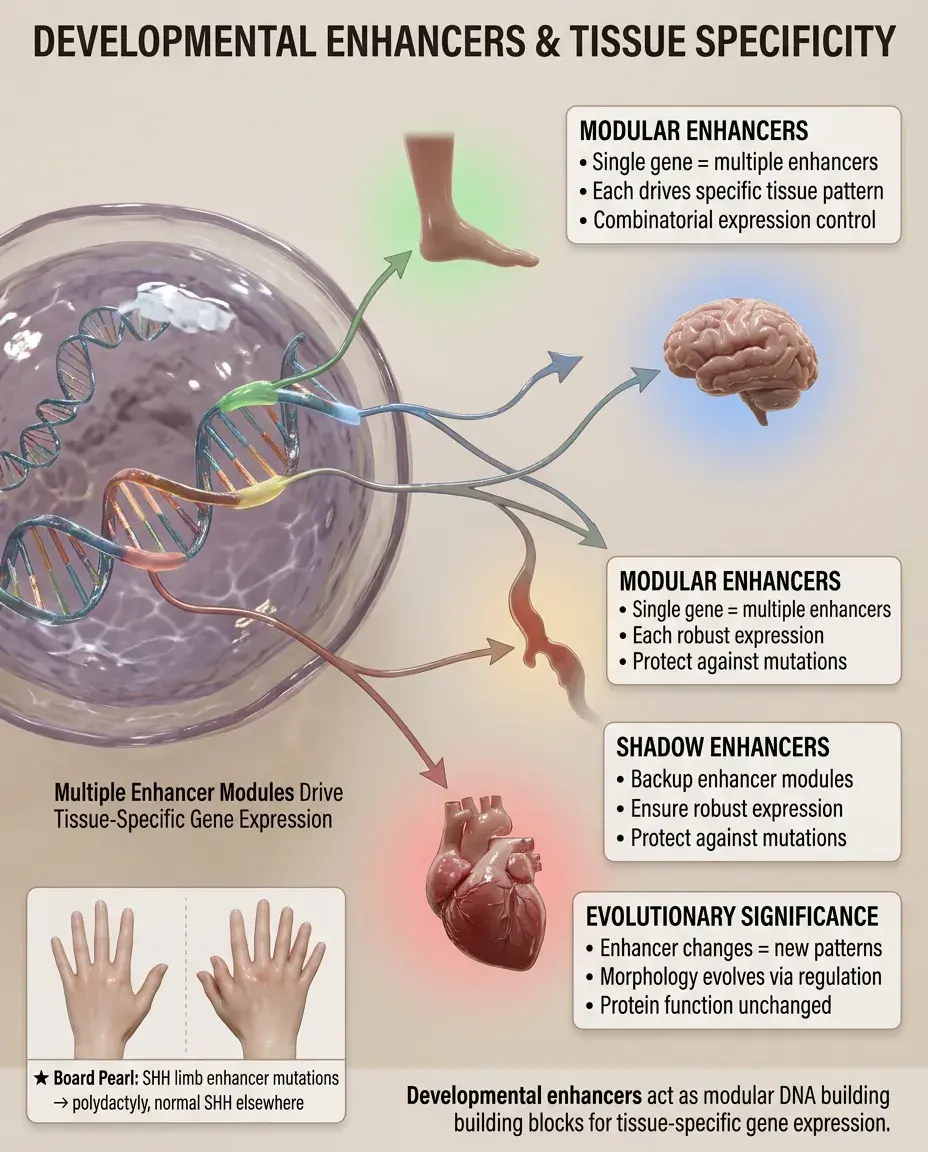
Developmental enhancers are DNA modules that drive gene expression in specific tissues, cell types, or developmental stages.
🧷
A single gene often has multiple enhancers creating its complete expression pattern — like modular building blocks.
🧷
Shadow enhancers are seemingly redundant enhancers that ensure robust expression despite environmental or genetic perturbations.
🧷
Enhancer evolution drives morphological evolution — changes in enhancers alter expression patterns without changing protein function.
🧷
Board pearl: Limb-specific enhancers of SHH control digit number — mutations cause polydactyly without affecting SHH expression elsewhere.
Chromatin States and Accessibility
📍
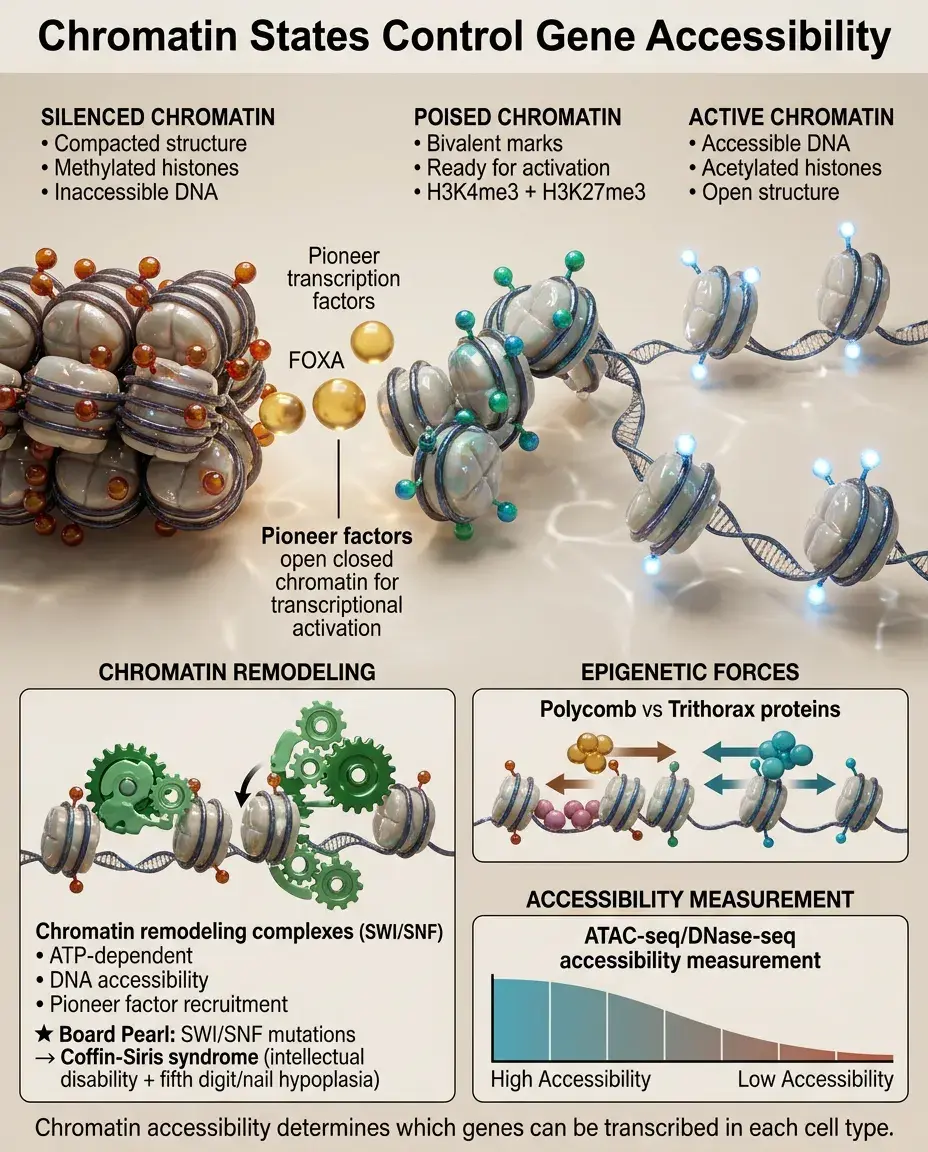
Developmental genes exist in different chromatin states: active (accessible, acetylated histones), poised (bivalent marks, ready for activation), or silenced (compacted, methylated).
📍
Pioneer factors like FOXA can bind to closed chromatin and recruit chromatin remodelers to make DNA accessible for other TFs.
📍
Chromatin accessibility is measured by techniques like ATAC-seq and DNase-seq, revealing which enhancers are active in each cell type.
📍
Polycomb and Trithorax group proteins maintain repressed and active states, respectively, through cell divisions.
📍
Board pearl: SWI/SNF chromatin remodeling complex mutations cause Coffin-Siris syndrome — intellectual disability with fifth digit/nail hypoplasia.
Transcriptional Memory and Cellular Inheritance
🔹
Transcriptional memory allows cells to remember past developmental decisions through multiple cell divisions.
🔹
Mechanisms include: DNA methylation patterns maintained by DNMT1, histone modifications propagated during replication, and positive feedback loops of TF expression.
🔹
Cellular memory is essential for maintaining cell type identity — a hepatocyte remains a hepatocyte through many divisions.
🔹
Memory can be erased during reprogramming (making iPSCs) or gametogenesis (epigenetic marks reset between generations).
🔹
Board pearl: Loss of transcriptional memory contributes to cancer — cells forget their differentiated state and reacquire stem-like properties.

Gene Regulatory Networks in Development
⭐
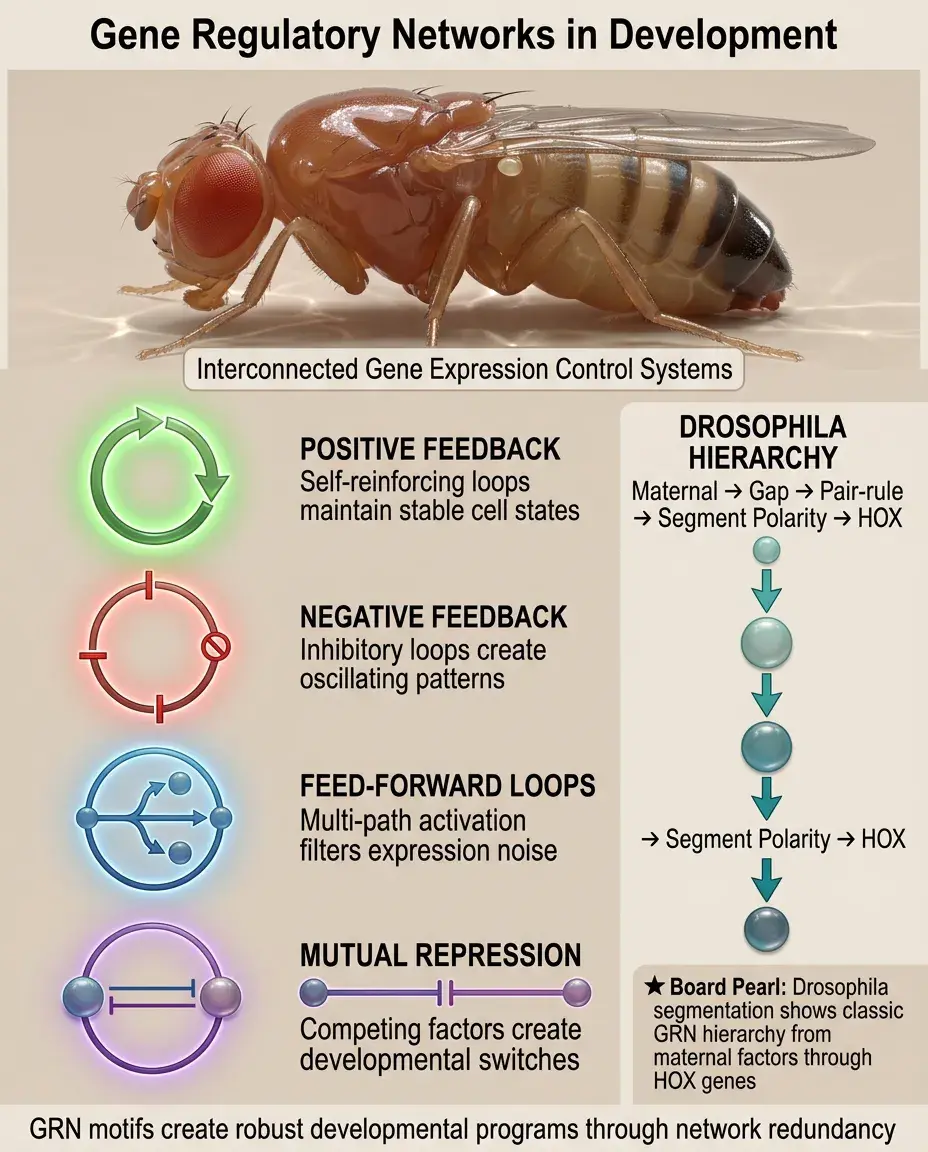
Development proceeds through gene regulatory networks (GRNs) — interconnected sets of genes that regulate each other's expression.
⭐
GRN motifs include: positive feedback (maintains cell state), negative feedback (creates oscillations), feed-forward loops (filters noise), and mutual repression (creates switches).
⭐
Robustness emerges from network properties — multiple pathways to the same outcome protect against perturbations.
⭐
Mathematical modeling of GRNs predicts how genetic changes alter developmental outcomes.
⭐
Board pearl: Segmentation in Drosophila demonstrates GRN hierarchy: maternal factors → gap genes → pair-rule genes → segment polarity genes → HOX genes.
Nuclear Organization and Gene Expression
✅
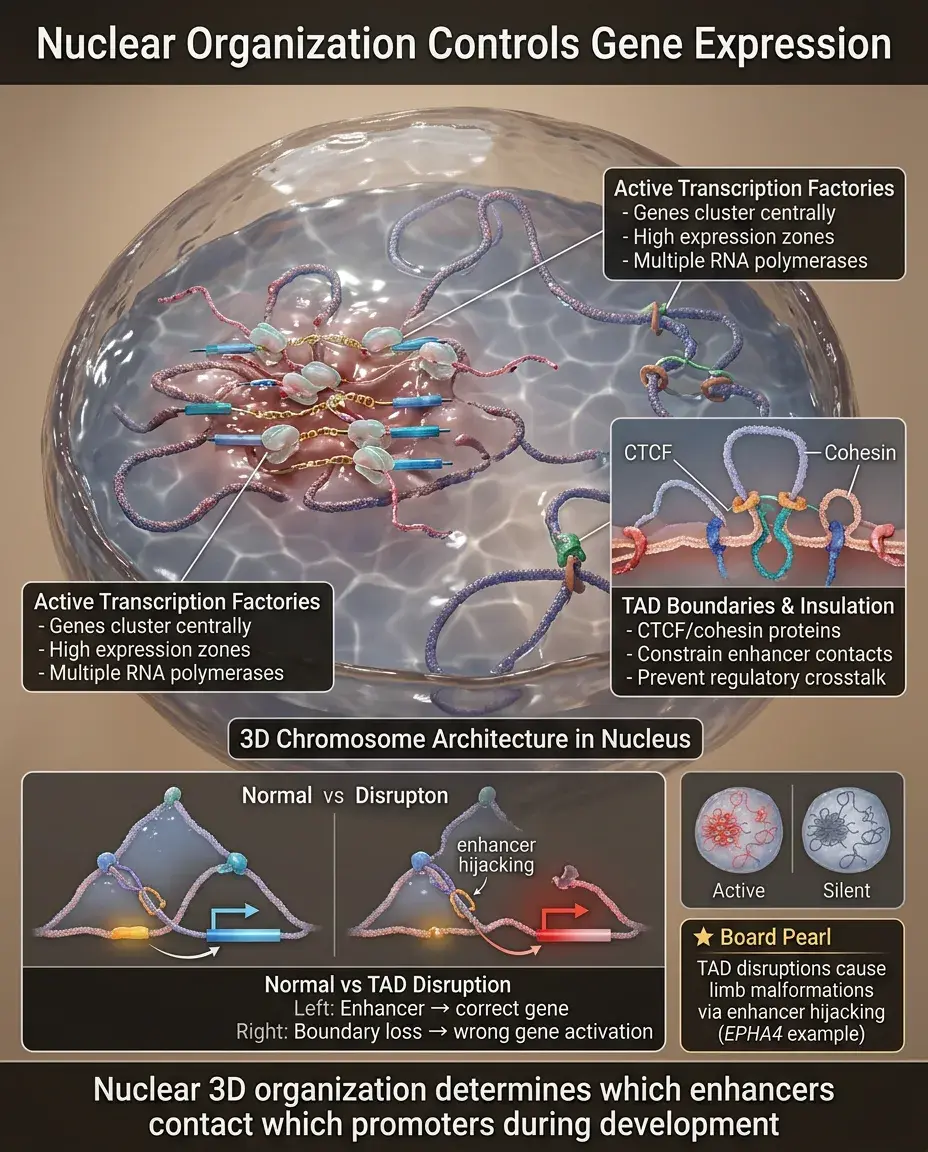
The 3D organization of chromosomes in the nucleus affects gene expression during development.
✅
Active genes cluster in transcription factories, while silent genes localize to the nuclear periphery or heterochromatic foci.
✅
TADs (topologically associating domains) are chromosomal regions that preferentially interact with each other, constraining enhancer-promoter contacts.
✅
CTCF and cohesin proteins create boundaries between TADs, insulating genes from inappropriate regulatory elements.
✅
Board pearl: TAD disruptions cause developmental disorders by allowing enhancers to activate wrong genes (e.g., limb malformations from EPHA4 enhancer hijacking).
Evolution of Developmental Gene Regulation
🧠
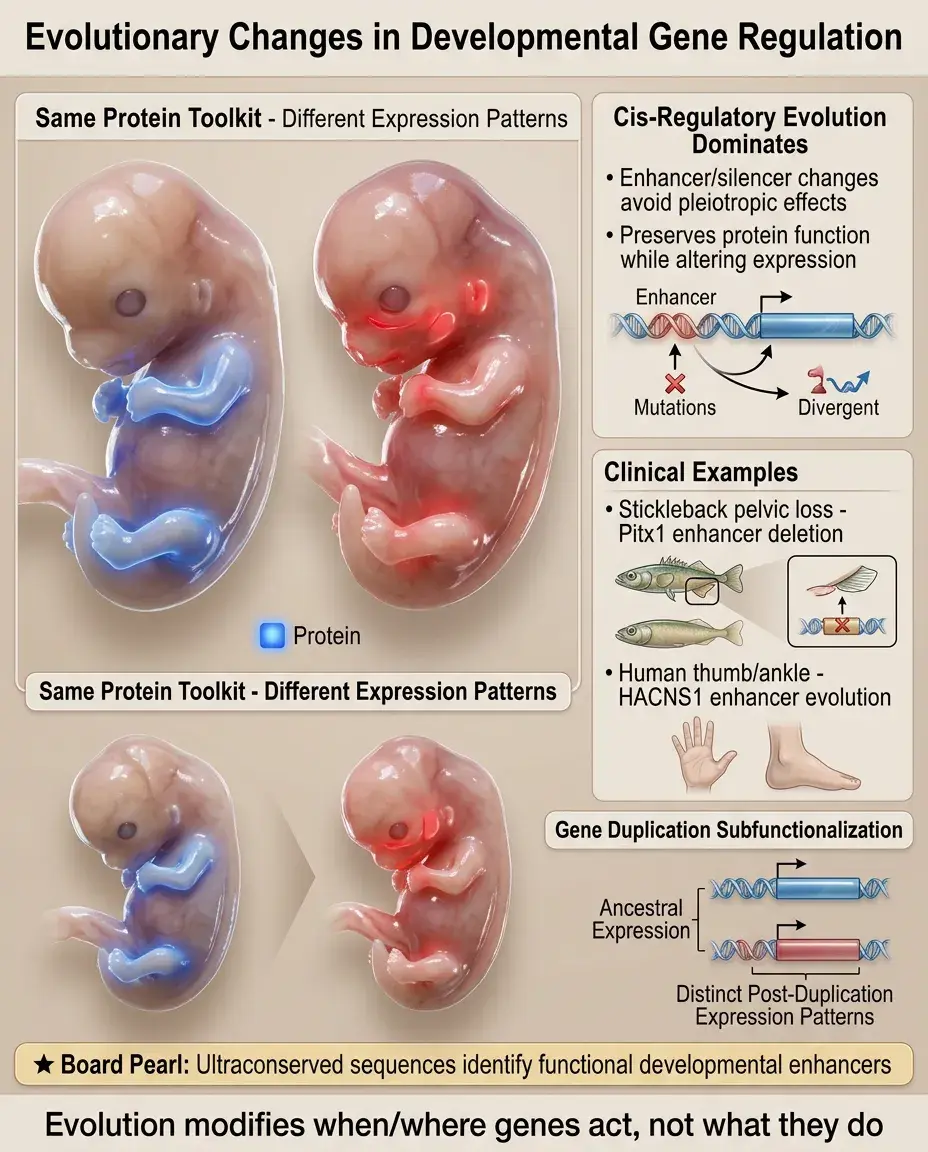
Evolution acts primarily on regulatory elements rather than protein-coding sequences — same toolkit, different expression patterns.
🧠
Cis-regulatory changes (affecting enhancers/silencers) predominate because they avoid pleiotropic effects of protein changes.
🧠
Examples: Stickleback pelvic loss (Pitx1 enhancer deletion), human-specific HACNS1 enhancer (thumb and ankle modifications).
🧠
Gene duplication allows subfunctionalization — duplicated genes partition ancestral expression patterns via enhancer divergence.
🧠
Board pearl: Evolutionary conservation identifies functionally important sequences — ultraconserved elements often contain developmental enhancers.
Transcriptional Noise and Precision
⚡
Gene expression is inherently stochastic — random bursts of transcription create cell-to-cell variability even in identical environments.
⚡
Developmental systems use multiple mechanisms to achieve precision despite noise: multiple enhancers, negative feedback, and cell-cell communication.
⚡
Some variability is beneficial — allows bet-hedging strategies and can be harnessed for stochastic cell fate decisions.
⚡
Single-cell RNA sequencing reveals heterogeneity in developing tissues previously thought to be homogeneous.
⚡
Board pearl: Haploinsufficiency disorders show variable expressivity partly due to transcriptional noise — borderline gene dosage plus stochastic variation.

Developmental Disorders from Regulatory Mutations
📌
Many developmental disorders result from mutations in regulatory elements rather than coding sequences.
📌
Enhanceropathies: diseases caused by enhancer mutations (e.g., Pierre Robin sequence from SOX9 enhancer deletions).
📌
Cohesinopathies: diseases from cohesin complex mutations affecting 3D genome organization (Cornelia de Lange syndrome).
📌
Laminopathies: diseases from nuclear lamina defects affecting heterochromatin organization (Hutchinson-Gilford progeria).
📌
Board distinction: Coding mutations often cause complete loss of function; regulatory mutations typically cause tissue-specific or partial loss.

Board Question Stem Patterns
📣
Mutation causes transformation of one body segment into another → homeotic gene mutation, likely HOX cluster.
📣
Multiple congenital anomalies from a single gene mutation → likely transcription factor controlling multiple developmental programs.
📣
Limb-specific defect with normal gene coding sequence → enhancer mutation affecting tissue-specific expression.
📣
Isotretinoin exposure causing specific pattern of defects → disruption of retinoic acid-regulated HOX gene expression.
📣
Variable expression of genetic disorder in family members → consider regulatory mutations with position effects or transcriptional noise.
📣
Cancer with chromosomal translocation creating fusion protein → developmental TF dysregulation (PAX3-FOXO1 in rhabdomyosarcoma).
📣
Syndrome with both skeletal and cognitive features → chromatin remodeler or general transcription machinery defect.

One-Line Recap
🔸
Developmental gene expression is controlled by transcription factors binding to enhancers and silencers, creating spatiotemporal patterns through combinatorial codes, morphogen gradients, and epigenetic modifications that establish and maintain cell fates via hierarchical gene regulatory networks whose disruption by coding or regulatory mutations causes developmental disorders and cancer.

bottom of page

